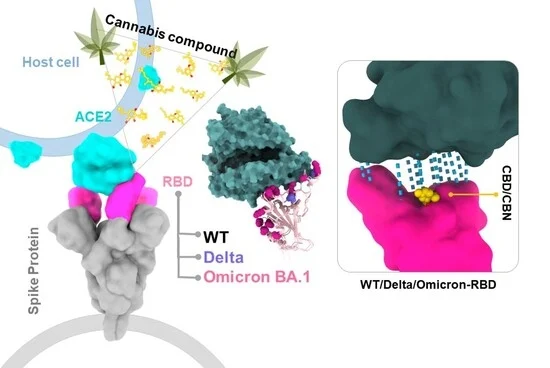
“Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) enters host cells when the spike receptor-binding domain (RBD) engages angiotensin-converting enzyme 2 (ACE2).
Cannabinoid scaffolds have recently been reported to bind S1/RBD, block spike-mediated membrane fusion, and modulate host inflammatory pathways, making them attractive candidates for entry inhibition.
Here, we applied an integrated computational pipeline to prioritize cannabis-derived compounds as interfacial blockers of the RBD-ACE2 complex across variants.
Eleven phytocannabinoids were docked into the wild-type (WT) RBD-ACE2 interface, identifying three cavities, with ligands preferentially occupying pocket 1. Complexes were subjected to triplicate 200 ns all-atom molecular dynamics (MD) simulations for WT, Delta, and Omicron BA.1 RBD-ACE2. Binding energetics were quantified using molecular mechanics/generalized Born surface area (MM/GBSA) and solvated interaction energy (SIE), and per-residue contributions were analyzed together with solvent-accessible surface area (SASA) and residue interaction networks.
Among all compounds, cannabidiol (CBD) and cannabinol (CBN) were the only ligands that remained stably bound in pocket 1 for all variants. CBN showed the most favorable ligand-complex binding in WT, whereas CBD preserved favorable binding in Omicron BA.1 despite reduced interface burial, indicating that van der Waals/electrostatic complementarity and solvation, rather than surface coverage alone, govern affinity. Both ligands weakened modeled RBD-ACE2 binding by perturbing hot-spot residues centered on Y505 or N501Y in RBD and E37, A387, and R393 in ACE2.
Overall, our results highlight CBD and CBN as tractable, variant-spanning interface disruptors and illustrate how MD-based free-energy calculations can support computational drug discovery against evolving viral protein-protein interfaces.”
https://pubmed.ncbi.nlm.nih.gov/42075932
“Together, these docking results support the hypothesis that specific phytocannabinoids can effectively occupy critical interface pockets of the RBD/ACE2 complex, with potential to sterically hinder or allosterically modulate spike–receptor recognition.”
https://www.mdpi.com/1420-3049/31/8/1253